2.3. Uncertainties in the theoretical predictions#
In this tutorial, you will learn how to use the microscopic module to print out some microscopic predictions for thermodynamical properties.
Import the libraries that will be employed in this tutorial.
# Import numpy
import numpy as np
# Import matplotlib
import matplotlib.pyplot as plt
# Import nucleardatapy package
import nucleardatapy as nuda
%matplotlib inline
You can simply print out the properties of the nuda’s function that we will use:
# Explore the nucleardatapy module to find the correct attribute
print(dir(nuda.matter.setupMicro))
['__class__', '__delattr__', '__dict__', '__dir__', '__doc__', '__eq__', '__format__', '__ge__', '__getattribute__', '__getstate__', '__gt__', '__hash__', '__init__', '__init_subclass__', '__le__', '__lt__', '__module__', '__ne__', '__new__', '__reduce__', '__reduce_ex__', '__repr__', '__setattr__', '__sizeof__', '__str__', '__subclasshook__', '__weakref__', 'init_self', 'print_outputs']
Get the full list of models:
models, models_lower = nuda.matter.micro_models()
print('models:',models)
models: ['1981-VAR-AM-FP', '1998-VAR-AM-APR', '1998-VAR-AM-APR-fit', '2012-AFDMC-NM-RES-1', '2012-AFDMC-NM-RES-2', '2012-AFDMC-NM-RES-3', '2012-AFDMC-NM-RES-4', '2012-AFDMC-NM-RES-5', '2012-AFDMC-NM-RES-6', '2012-AFDMC-NM-RES-7', '2012-AFDMC-NM-FIT-1', '2012-AFDMC-NM-FIT-2', '2012-AFDMC-NM-FIT-3', '2012-AFDMC-NM-FIT-4', '2012-AFDMC-NM-FIT-5', '2012-AFDMC-NM-FIT-6', '2012-AFDMC-NM-FIT-7', '2022-AFDMC-NM', '2024-BHF-AM-2BF-Av18', '2024-BHF-AM-2BF-BONN', '2024-BHF-AM-2BF-CDBONN', '2024-BHF-AM-2BF-NSC97a', '2024-BHF-AM-2BF-NSC97b', '2024-BHF-AM-2BF-NSC97c', '2024-BHF-AM-2BF-NSC97d', '2024-BHF-AM-2BF-NSC97e', '2024-BHF-AM-2BF-NSC97f', '2006-BHF-AM-Av18', '2024-BHF-AM-23BF-Av18', '2024-BHF-AM-23BF-BONN', '2024-BHF-AM-23BF-CDBONN', '2024-BHF-AM-23BF-NSC97a', '2024-BHF-AM-23BF-NSC97b', '2024-BHF-AM-23BF-NSC97c', '2024-BHF-AM-23BF-NSC97d', '2024-BHF-AM-23BF-NSC97e', '2024-BHF-AM-23BF-NSC97f', '2008-QMC-NM-swave', '2010-QMC-NM-AV4', '2009-DLQMC-NM', '2014-AFQMC-NM', '2016-QMC-NM', '2018-QMC-NM', '2024-QMC-NM', '2013-MBPT-NM', '2016-MBPT-AM', '2019-MBPT-AM-L59', '2019-MBPT-AM-L69', '2020-MBPT-AM', '2024-MBPT-AM-DN2LO-450', '2024-MBPT-AM-DN2LO-500', '2024-MBPT-AM-DN2LOgo-394', '2024-MBPT-AM-DN2LOgo-450', '2024-MBPT-AM-N2LOsat', '2024-NLEFT-AM', '2020-SCGF-AM-N3LO-414', '2020-SCGF-AM-N3LO-450', '2020-SCGF-AM-N3LO-500', '2024-SCGF-AM-DN2LO-450', '2024-SCGF-AM-DN2LO-500', '2024-SCGF-AM-DN2LOgo-394', '2024-SCGF-AM-DN2LOgo-450', '2024-SCGF-AM-N2LOsat', '2024-CC-AM-DN2LO-450', '2024-CC-AM-DN2LO-500', '2024-CC-AM-DN2LOgo-394', '2024-CC-AM-DN2LOgo-450', '2024-CC-AM-N2LOsat']
Fix one model for which we want to extract the data:
model = '2018-QMC-NM'
mic = nuda.matter.setupMicro( model = model )
model -> 2018-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
print the outputs:
mic.print_outputs()
- Print output:
model: 2018-QMC-NM
ref: I. Tews, J. Carlson, S. Gandolfi, S. Reddy, Astroph. J. 860(2), 149 (2018).
label: QMC-2018
note:
self.sm_den: None
self.sm_effmass: None
den: [0.01 0.031 0.051 0.072 0.092 0.113 0.133 0.154 0.174 0.195 0.215 0.236
0.256 0.277 0.297 0.318 0.338 0.359 0.379 0.4 ] in fm$^{-3}$
kfn: [0.01 0.031 0.051 0.072 0.092 0.113 0.133 0.154 0.174 0.195 0.215 0.236
0.256 0.277 0.297 0.318 0.338 0.359 0.379 0.4 ] in fm$^{-1}$
asy: 0.0
nm_den: [0.02 0.04 0.06 0.08 0.1 0.12 0.14 0.16 0.18 0.2 0.22 0.24 0.26 0.28
0.3 0.32] in fm$^{-3}$
nm_kfn: [0.84 1.058 1.211 1.333 1.436 1.526 1.606 1.68 1.747 1.809 1.868 1.923
1.975 2.024 2.071 2.116] in fm$^{-1}$
nm_e2a: [944.084 946.293 948.039 949.658 951.231 952.797 954.406 956.061 957.794
959.592 961.46 963.39 965.406 967.425 969.633 971.778] in MeV
nm_e2a_err: [ 0.071 0.118 0.265 0.499 0.815 1.213 1.717 2.307 3.023 3.828
4.741 5.78 6.944 8.227 9.662 11.134] in MeV
nm_eps: [ 18.882 37.852 56.882 75.973 95.123 114.336 133.617 152.97 172.403
191.918 211.521 231.214 251.005 270.879 290.89 310.969] in MeV fm$^{-3}$
nm_eps_err: [1.000e-03 5.000e-03 1.600e-02 4.000e-02 8.100e-02 1.460e-01 2.400e-01
3.690e-01 5.440e-01 7.660e-01 1.043e+00 1.387e+00 1.805e+00 2.304e+00
2.899e+00 3.563e+00] in MeV fm$^{-3}$
nm_pre: [ 0.053 0.152 0.298 0.509 0.78 1.141 1.595 2.167 2.862 3.67
4.578 5.727 6.753 8.288 10.058 10.27 ] in MeV fm$^{-3}$
nm_cs2: [ 0.004 0.006 0.009 0.012 0.016 0.021 0.026 0.033 0.039 0.042
0.054 0.054 0.06 0.093 0.067 -0.063]
The uncertianty is defined as:
print('uncertainties:',mic.nm_e2a_err/mic.nm_e2a)
uncertainties: [7.52051449e-05 1.24327198e-04 2.79935751e-04 5.25557873e-04
8.56469462e-04 1.27288455e-03 1.79902540e-03 2.41271280e-03
3.15641938e-03 3.98929776e-03 4.93120071e-03 5.99995944e-03
7.19257165e-03 8.50381566e-03 9.96459952e-03 1.14573441e-02]
Fix a list of models instead of a unique one
models = [ '2018-QMC-NM', '2016-MBPT-AM', '2016-QMC-NM', '2020-MBPT-AM', '2024-QMC-NM' ]
print('models:',models)
for model in models:
print('model:',model)
mic = nuda.matter.setupMicro( model = model )
models: ['2018-QMC-NM', '2016-MBPT-AM', '2016-QMC-NM', '2020-MBPT-AM', '2024-QMC-NM']
model: 2018-QMC-NM
model -> 2018-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model: 2016-MBPT-AM
model -> 2016-MBPT-AM
flag_nm: True
flag_sm: True
flag_kf: False
flag_den: True
model: 2016-QMC-NM
model -> 2016-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model: 2020-MBPT-AM
model -> 2020-MBPT-AM
flag_nm: True
flag_sm: True
flag_kf: False
flag_den: True
model: 2024-QMC-NM
model -> 2024-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: False
flag_den: True
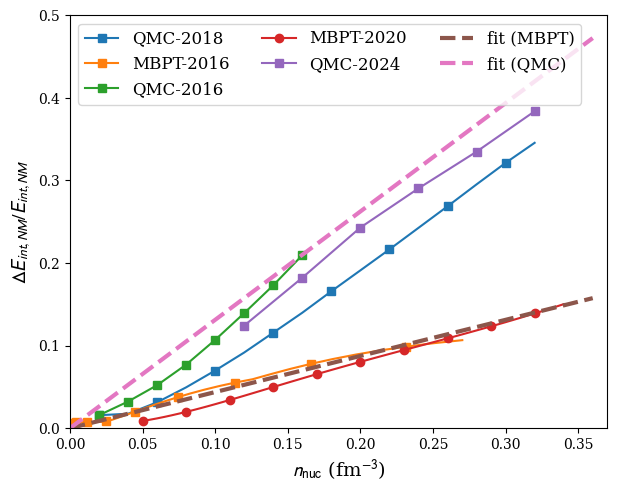
plot:
nuda.fig.matter_setupMicro_err_NM_fig( None, models )
Plot name: None
model -> 2018-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model: 2018-QMC-NM
model -> 2016-MBPT-AM
flag_nm: True
flag_sm: True
flag_kf: False
flag_den: True
model: 2016-MBPT-AM
model -> 2016-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model: 2016-QMC-NM
model -> 2020-MBPT-AM
flag_nm: True
flag_sm: True
flag_kf: False
flag_den: True
model: 2020-MBPT-AM
model -> 2024-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: False
flag_den: True
model: 2024-QMC-NM
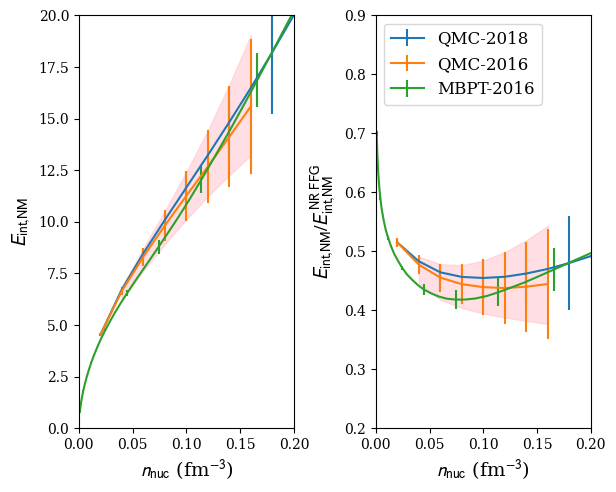
We now define a reference band in NM:
We first choose as set of calculations existing in nuda’s library. We first need to know what are the calculations available:
models, models_lower = nuda.matter.micro_models()
print('models:',models)
models: ['1981-VAR-AM-FP', '1998-VAR-AM-APR', '1998-VAR-AM-APR-fit', '2012-AFDMC-NM-RES-1', '2012-AFDMC-NM-RES-2', '2012-AFDMC-NM-RES-3', '2012-AFDMC-NM-RES-4', '2012-AFDMC-NM-RES-5', '2012-AFDMC-NM-RES-6', '2012-AFDMC-NM-RES-7', '2012-AFDMC-NM-FIT-1', '2012-AFDMC-NM-FIT-2', '2012-AFDMC-NM-FIT-3', '2012-AFDMC-NM-FIT-4', '2012-AFDMC-NM-FIT-5', '2012-AFDMC-NM-FIT-6', '2012-AFDMC-NM-FIT-7', '2022-AFDMC-NM', '2024-BHF-AM-2BF-Av18', '2024-BHF-AM-2BF-BONN', '2024-BHF-AM-2BF-CDBONN', '2024-BHF-AM-2BF-NSC97a', '2024-BHF-AM-2BF-NSC97b', '2024-BHF-AM-2BF-NSC97c', '2024-BHF-AM-2BF-NSC97d', '2024-BHF-AM-2BF-NSC97e', '2024-BHF-AM-2BF-NSC97f', '2006-BHF-AM-Av18', '2024-BHF-AM-23BF-Av18', '2024-BHF-AM-23BF-BONN', '2024-BHF-AM-23BF-CDBONN', '2024-BHF-AM-23BF-NSC97a', '2024-BHF-AM-23BF-NSC97b', '2024-BHF-AM-23BF-NSC97c', '2024-BHF-AM-23BF-NSC97d', '2024-BHF-AM-23BF-NSC97e', '2024-BHF-AM-23BF-NSC97f', '2008-QMC-NM-swave', '2010-QMC-NM-AV4', '2009-DLQMC-NM', '2014-AFQMC-NM', '2016-QMC-NM', '2018-QMC-NM', '2024-QMC-NM', '2013-MBPT-NM', '2016-MBPT-AM', '2019-MBPT-AM-L59', '2019-MBPT-AM-L69', '2020-MBPT-AM', '2024-MBPT-AM-DN2LO-450', '2024-MBPT-AM-DN2LO-500', '2024-MBPT-AM-DN2LOgo-394', '2024-MBPT-AM-DN2LOgo-450', '2024-MBPT-AM-N2LOsat', '2024-NLEFT-AM', '2020-SCGF-AM-N3LO-414', '2020-SCGF-AM-N3LO-450', '2020-SCGF-AM-N3LO-500', '2024-SCGF-AM-DN2LO-450', '2024-SCGF-AM-DN2LO-500', '2024-SCGF-AM-DN2LOgo-394', '2024-SCGF-AM-DN2LOgo-450', '2024-SCGF-AM-N2LOsat', '2024-CC-AM-DN2LO-450', '2024-CC-AM-DN2LO-500', '2024-CC-AM-DN2LOgo-394', '2024-CC-AM-DN2LOgo-450', '2024-CC-AM-N2LOsat']
Then we select a few of them:
models = [ '2018-QMC-NM', '2016-QMC-NM', '2016-MBPT-AM' ]
The reference band in NM is then calculated in the following way:
band = nuda.matter.setupMicroBand( models, matter='NM' )
band.print_outputs( )
Band in NM
model -> 2018-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model -> 2016-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model -> 2016-MBPT-AM
flag_nm: True
flag_sm: True
flag_kf: False
flag_den: True
model -> 2018-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model -> 2016-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model -> 2016-MBPT-AM
flag_nm: True
flag_sm: True
flag_kf: False
flag_den: True
model -> 2018-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model -> 2016-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model -> 2016-MBPT-AM
flag_nm: True
flag_sm: True
flag_kf: False
flag_den: True
- Print output:
models : ['2018-QMC-NM', '2016-QMC-NM', '2016-MBPT-AM']
den_min: 0.02
den_max: 0.16
den : [0.02 0.034 0.048 0.062 0.076 0.09 0.104 0.118 0.132 0.146 0.16 ]
kfn : [0.84 1. 1.12 1.22 1.31 1.39 1.45 1.52 1.58 1.63 1.68]
e2a : [ 4.5 5.95 7.22 8.33 9.42 10.5 11.58 12.68 13.8 14.96 16.14]
std : [0.017 0.37 0.434 0.572 0.745 0.965 1.232 1.553 1.941 2.397 2.918]
Plot:
den = np.linspace(0.04,0.16,num=7)
print('den:',den)
nuda.fig.matter_setupMicro_band_fig( None, models, den=den, matter='NM' )
den: [0.04 0.06 0.08 0.1 0.12 0.14 0.16]
Plot name: None
model -> 2018-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model: 2018-QMC-NM
model -> 2016-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model: 2016-QMC-NM
model -> 2016-MBPT-AM
flag_nm: True
flag_sm: True
flag_kf: False
flag_den: True
model: 2016-MBPT-AM
Band in NM
model -> 2018-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model -> 2016-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model -> 2016-MBPT-AM
flag_nm: True
flag_sm: True
flag_kf: False
flag_den: True
model -> 2018-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model -> 2016-QMC-NM
flag_nm: True
flag_sm: False
flag_kf: True
flag_den: False
model -> 2016-MBPT-AM
flag_nm: True
flag_sm: True
flag_kf: False
flag_den: True