2.5. Pairing gap (micro)#
In this tutorial, you will learn how to employ the pairing gap module.
Import the libraries that will be employed in this tutorial.
# Import numpy
import numpy as np
# Import matplotlib
import matplotlib.pyplot as plt
# Import nucleardatapy package
import nucleardatapy as nuda
%matplotlib inline
You can simply print out the properties of the nuda’s function that we will use:
# Explore the nucleardatapy module to find the correct attribute
print(dir(nuda.matter.setupMicroGap))
['__class__', '__delattr__', '__dict__', '__dir__', '__doc__', '__eq__', '__format__', '__ge__', '__getattribute__', '__getstate__', '__gt__', '__hash__', '__init__', '__init_subclass__', '__le__', '__lt__', '__module__', '__ne__', '__new__', '__reduce__', '__reduce_ex__', '__repr__', '__setattr__', '__sizeof__', '__str__', '__subclasshook__', '__weakref__', 'init_self', 'print_outputs']
Get the full list of models ( matter can be ‘NM’ (default) or ‘SM’ ) :
models, models_lower, models_all, models_all_lower = nuda.matter.micro_gap_models( matter = 'NM' )
print('models:',models)
models: ['2006-BHF-NM-Av18', '2008-BCS-NM', '2008-QMC-NM-swave', '2009-DLQMC-NM', '2010-QMC-NM-AV4', '2017-MBPT-NM-GAP-EMG-450-500-N2LO', '2017-MBPT-NM-GAP-EMG-450-500-N3LO', '2017-MBPT-NM-GAP-EMG-450-700-N2LO', '2017-MBPT-NM-GAP-EMG-450-700-N3LO', '2017-MBPT-NM-GAP-EM-500-N2LO', '2017-MBPT-NM-GAP-EM-500-N3LO', '2022-AFDMC-NM']
Fix one model for which we want to extract the data:
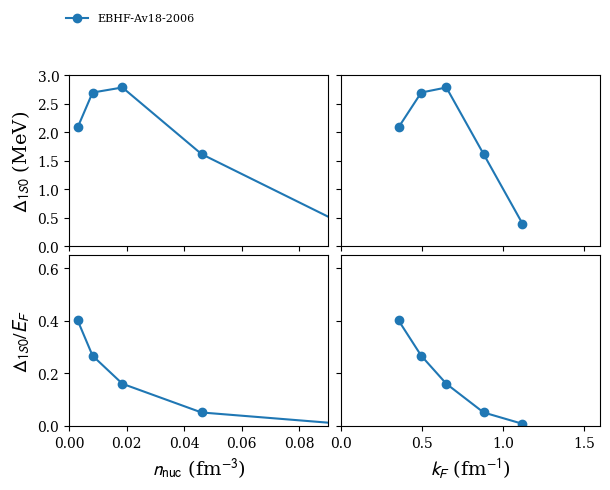
model = '2006-BHF-NM-Av18'
Instantiate the object gap with the pairing gap data:
pheno = nuda.matter.setupMicroGap( model = model )
pheno.print_outputs()
-> model: 2006-BHF-NM-Av18
- Print output:
model: 2006-BHF-NM-Av18
ref: L.G. Cao, U. Lombardo, and P. Schuck, Phys. Rev. C 74, 64301 (2006)
note:
label: EBHF-Av18-2006
marker: o
every: 1
nm_den_1s0: [0. 0.002 0.008 0.019 0.037 0.064 0.081] in fm$^{-3}$
nm_kfn_1s0: [0.186 0.385 0.608 0.822 1.03 1.24 1.34 ] in fm$^{-1}$
nm_gap_1s0: [0.37 1.04 1.55 1.86 1.54 0.47 0. ] in MeV
Fix a list of params instead of a unique one
models, models_lower, models_all, models_all_lower = nuda.matter.micro_gap_models( matter = 'NM' )
print('models:',models)
for model in models:
print('model:',model)
gap = nuda.matter.setupMicroGap( model = model )
models: ['2006-BHF-NM-Av18', '2008-BCS-NM', '2008-QMC-NM-swave', '2009-DLQMC-NM', '2010-QMC-NM-AV4', '2017-MBPT-NM-GAP-EMG-450-500-N2LO', '2017-MBPT-NM-GAP-EMG-450-500-N3LO', '2017-MBPT-NM-GAP-EMG-450-700-N2LO', '2017-MBPT-NM-GAP-EMG-450-700-N3LO', '2017-MBPT-NM-GAP-EM-500-N2LO', '2017-MBPT-NM-GAP-EM-500-N3LO', '2022-AFDMC-NM']
model: 2006-BHF-NM-Av18
-> model: 2006-BHF-NM-Av18
model: 2008-BCS-NM
-> model: 2008-BCS-NM
model: 2008-QMC-NM-swave
-> model: 2008-QMC-NM-swave
model: 2009-DLQMC-NM
-> model: 2009-DLQMC-NM
model: 2010-QMC-NM-AV4
-> model: 2010-QMC-NM-AV4
model: 2017-MBPT-NM-GAP-EMG-450-500-N2LO
-> model: 2017-MBPT-NM-GAP-EMG-450-500-N2LO
model: 2017-MBPT-NM-GAP-EMG-450-500-N3LO
-> model: 2017-MBPT-NM-GAP-EMG-450-500-N3LO
model: 2017-MBPT-NM-GAP-EMG-450-700-N2LO
-> model: 2017-MBPT-NM-GAP-EMG-450-700-N2LO
model: 2017-MBPT-NM-GAP-EMG-450-700-N3LO
-> model: 2017-MBPT-NM-GAP-EMG-450-700-N3LO
model: 2017-MBPT-NM-GAP-EM-500-N2LO
-> model: 2017-MBPT-NM-GAP-EM-500-N2LO
model: 2017-MBPT-NM-GAP-EM-500-N3LO
-> model: 2017-MBPT-NM-GAP-EM-500-N3LO
model: 2022-AFDMC-NM
-> model: 2022-AFDMC-NM
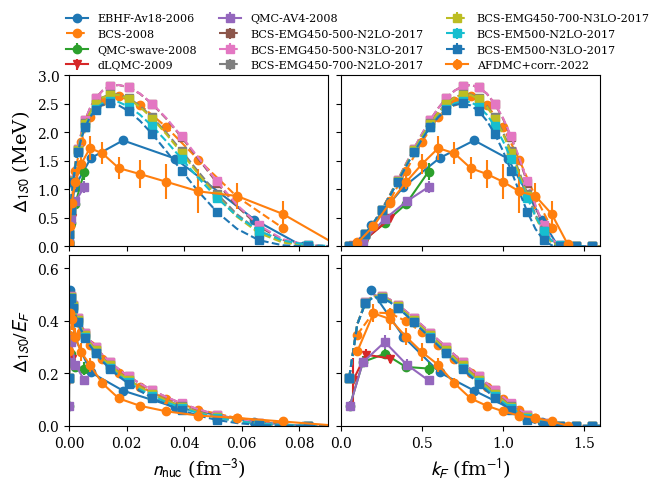
plot 1S0 pairing gap in NM:
nuda.fig.matter_setupMicro_gap_1s0_fig( None, models, matter = 'NM' )
Plot name: None
models: ['2006-BHF-NM-Av18', '2008-BCS-NM', '2008-QMC-NM-swave', '2009-DLQMC-NM', '2010-QMC-NM-AV4', '2017-MBPT-NM-GAP-EMG-450-500-N2LO', '2017-MBPT-NM-GAP-EMG-450-500-N3LO', '2017-MBPT-NM-GAP-EMG-450-700-N2LO', '2017-MBPT-NM-GAP-EMG-450-700-N3LO', '2017-MBPT-NM-GAP-EM-500-N2LO', '2017-MBPT-NM-GAP-EM-500-N3LO', '2022-AFDMC-NM']
-> model: 2006-BHF-NM-Av18
-> model: 2008-BCS-NM
-> model: 2008-QMC-NM-swave
-> model: 2009-DLQMC-NM
-> model: 2010-QMC-NM-AV4
-> model: 2017-MBPT-NM-GAP-EMG-450-500-N2LO
-> model: 2017-MBPT-NM-GAP-EMG-450-500-N3LO
-> model: 2017-MBPT-NM-GAP-EMG-450-700-N2LO
-> model: 2017-MBPT-NM-GAP-EMG-450-700-N3LO
-> model: 2017-MBPT-NM-GAP-EM-500-N2LO
-> model: 2017-MBPT-NM-GAP-EM-500-N3LO
-> model: 2022-AFDMC-NM
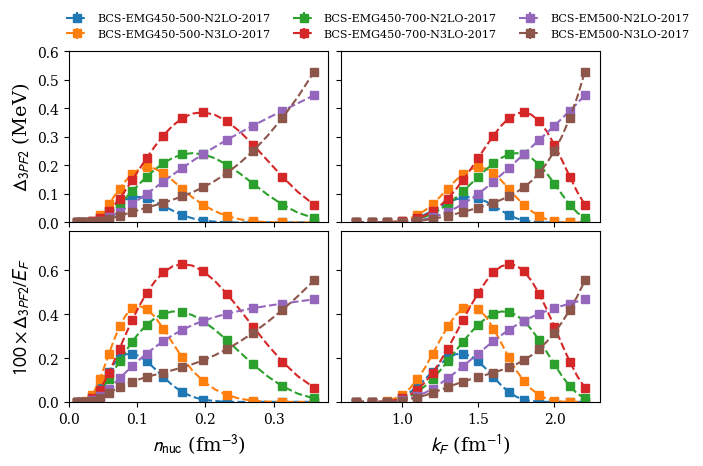
plot 3PGF2 pairing gap in NM:
nuda.fig.matter_setupMicro_gap_3pf2_fig( None, models, matter = 'NM' )
Plot name: None
models: ['2006-BHF-NM-Av18', '2008-BCS-NM', '2008-QMC-NM-swave', '2009-DLQMC-NM', '2010-QMC-NM-AV4', '2017-MBPT-NM-GAP-EMG-450-500-N2LO', '2017-MBPT-NM-GAP-EMG-450-500-N3LO', '2017-MBPT-NM-GAP-EMG-450-700-N2LO', '2017-MBPT-NM-GAP-EMG-450-700-N3LO', '2017-MBPT-NM-GAP-EM-500-N2LO', '2017-MBPT-NM-GAP-EM-500-N3LO', '2022-AFDMC-NM']
-> model: 2006-BHF-NM-Av18
-> model: 2008-BCS-NM
-> model: 2008-QMC-NM-swave
-> model: 2009-DLQMC-NM
-> model: 2010-QMC-NM-AV4
-> model: 2017-MBPT-NM-GAP-EMG-450-500-N2LO
-> model: 2017-MBPT-NM-GAP-EMG-450-500-N3LO
-> model: 2017-MBPT-NM-GAP-EMG-450-700-N2LO
-> model: 2017-MBPT-NM-GAP-EMG-450-700-N3LO
-> model: 2017-MBPT-NM-GAP-EM-500-N2LO
-> model: 2017-MBPT-NM-GAP-EM-500-N3LO
-> model: 2022-AFDMC-NM
plot 1S0 pairing gap in SM:
models, models_lower, models_all, models_all_lower = nuda.matter.micro_gap_models( matter = 'SM' )
print('models:',models)
nuda.fig.matter_setupMicro_gap_1s0_fig( None, models, matter = 'SM' )
models: ['2006-BHF-SM-Av18']
Plot name: None
models: ['2006-BHF-SM-Av18']
-> model: 2006-BHF-SM-Av18