6.1. Neutron Star Crust#
In this tutorial, you will learn how to extract neutron star crust EoS.
Import the libraries that will be employed in this tutorial.
# Import numpy
import numpy as np
# Import matplotlib
import matplotlib.pyplot as plt
# Import nucleardatapy package
import nucleardatapy as nuda
%matplotlib inline
You can simply print out the properties of the nuda’s function that we will use:
# Explore the nucleardatapy module to find the correct attribute
print(dir(nuda.crust.setupCrust))
['__class__', '__delattr__', '__dict__', '__dir__', '__doc__', '__eq__', '__format__', '__ge__', '__getattribute__', '__getstate__', '__gt__', '__hash__', '__init__', '__init_subclass__', '__le__', '__lt__', '__module__', '__ne__', '__new__', '__reduce__', '__reduce_ex__', '__repr__', '__setattr__', '__sizeof__', '__str__', '__subclasshook__', '__weakref__', 'init_self', 'print_outputs']
Get the full list of models:
models, models_lower = nuda.crust.crust_models()
print('models:',models)
models: ['1973-Negele-Vautherin', '2018-PCPFDDG-BSK22', '2018-PCPFDDG-BSK24', '2018-PCPFDDG-BSK25', '2018-PCPFDDG-BSK26', '2020-MVCD-D1S', '2020-MVCD-D1M', '2020-MVCD-D1MS', '2022-GMRS-BSK14', '2022-GMRS-BSK16', '2022-GMRS-DHSL59', '2022-GMRS-DHSL69', '2022-GMRS-F0', '2022-GMRS-H1', '2022-GMRS-H2', '2022-GMRS-H3', '2022-GMRS-H4', '2022-GMRS-H5', '2022-GMRS-H7', '2022-GMRS-LNS5', '2022-GMRS-RATP', '2022-GMRS-SGII', '2022-GMRS-SLY5']
for model in models:
crust = nuda.crust.setupCrust( model = model )
#crust.print_outputs( )
---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
Cell In[4], line 2
1 for model in models:
----> 2 crust = nuda.crust.setupCrust( model = model )
3 #crust.print_outputs( )
File ~/work/nucleardatapy/nucleardatapy/version-1.1/nucleardatapy/crust/setup_crust.py:250, in setupCrust.__init__(self, model)
245 self.latexCite = 'MPearson:2018'
246 self.ncl = 0.16 # in fm-3
247 self.den, self.Z, self.xp, self.N, A, self.RWS, xlambda, self.pre, self.e2a_etf, self.e2a_tot, \
248 self.rho, siswitch, self.e_si, self.e_pair, self.mu_p, self.mu_n, self.mu_e, ap, Cp, an, Cp, nLP, nLn, \
249 nBp, nBn, self.Zcl, self.Ncl, self.Acl \
--> 250 = np.loadtxt( file_in, usecols=(0,1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28), unpack = True )
251 #self.den, self.Z, self.xp, self.N, self.RWS, self.pre, self.e2a_etf, self.e2a_int, self.e2a_tot, self.mu_p, self.mu_n, self.mu_e, self.den_f,self.Zcl, self.Ncl \
252 # = np.loadtxt( file_in, usecols=(0,1,2,3,5,7,8,9,10,14,15,16,23,25,27), unpack = True )
253 self.den_cgs = self.den / 1.e-39 # in cm-3
File /opt/hostedtoolcache/Python/3.12.13/x64/lib/python3.12/site-packages/numpy/lib/_npyio_impl.py:1384, in loadtxt(fname, dtype, comments, delimiter, converters, skiprows, usecols, unpack, ndmin, encoding, max_rows, quotechar, like)
1381 if isinstance(delimiter, bytes):
1382 delimiter = delimiter.decode('latin1')
-> 1384 arr = _read(fname, dtype=dtype, comment=comment, delimiter=delimiter,
1385 converters=converters, skiplines=skiprows, usecols=usecols,
1386 unpack=unpack, ndmin=ndmin, encoding=encoding,
1387 max_rows=max_rows, quote=quotechar)
1389 return arr
File /opt/hostedtoolcache/Python/3.12.13/x64/lib/python3.12/site-packages/numpy/lib/_npyio_impl.py:1035, in _read(fname, delimiter, comment, quote, imaginary_unit, usecols, skiplines, max_rows, converters, ndmin, unpack, dtype, encoding)
1032 data = _preprocess_comments(data, comments, encoding)
1034 if read_dtype_via_object_chunks is None:
-> 1035 arr = _load_from_filelike(
1036 data, delimiter=delimiter, comment=comment, quote=quote,
1037 imaginary_unit=imaginary_unit,
1038 usecols=usecols, skiplines=skiplines, max_rows=max_rows,
1039 converters=converters, dtype=dtype,
1040 encoding=encoding, filelike=filelike,
1041 byte_converters=byte_converters)
1043 else:
1044 # This branch reads the file into chunks of object arrays and then
1045 # casts them to the desired actual dtype. This ensures correct
1046 # string-length and datetime-unit discovery (like `arr.astype()`).
1047 # Due to chunking, certain error reports are less clear, currently.
1048 if filelike:
ValueError: invalid column index 28 at row 1 with 28 columns
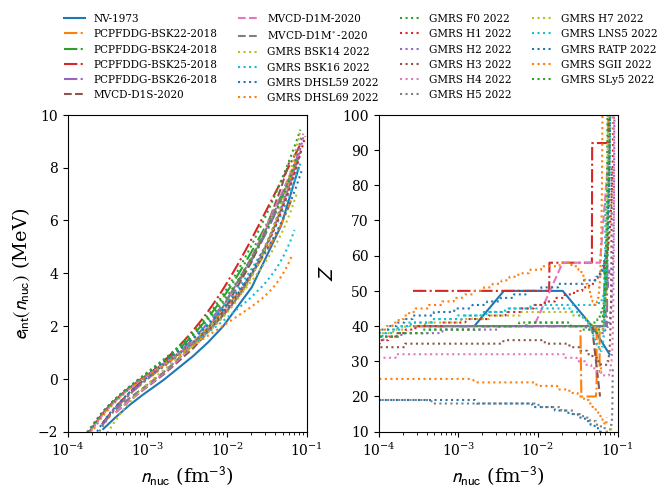
figure:
nuda.fig.crust_setupCrust_fig( None, models )
Plot name: None
model: 1973-Negele-Vautherin
model: 2018-PCPFDDG-BSK22
model: 2018-PCPFDDG-BSK24
model: 2018-PCPFDDG-BSK25
model: 2018-PCPFDDG-BSK26
model: 2020-MVCD-D1S
model: 2020-MVCD-D1M
model: 2020-MVCD-D1MS
model: 2022-GMRS-BSK14
model: 2022-GMRS-BSK16
model: 2022-GMRS-DHSL59
model: 2022-GMRS-DHSL69
model: 2022-GMRS-F0
model: 2022-GMRS-H1
model: 2022-GMRS-H2
model: 2022-GMRS-H3
model: 2022-GMRS-H4
model: 2022-GMRS-H5
model: 2022-GMRS-H7
model: 2022-GMRS-LNS5
model: 2022-GMRS-RATP
model: 2022-GMRS-SGII
model: 2022-GMRS-SLY5